Retroviridae
Introduction:
Vertebrate retroviruses (Retroviridae) are restricted to vertebrate animals. They are viral particles that reverse transcribe their RNA genome into a double stranded DNA copy that is inserted into the infected host cell genome. Their diploid RNA genome is enveloped within a protein capsid by a membrane fragment of the host cell in which env antigens are embedded.
Genomic structure:
Retroviridae originally received attention when infectious representatives were characterized in humans. However, it is now known that any LTR retrotransposon capable of recruiting a third ORF envelope gene (env) is potentially capable of becoming a retrovirus (env is the most basic difference between LTR retrotransposons and retroviruses). In fact, the Retroviridae display a gag-pol structure similar to that presented by Ty3/Gypsy LTR retroelements; the absence or presence of an env gene is the main difference between a Ty3/Gypsy LTR retrotransposon and a potential Ty3/Gypsy or Retroviridae simple retrovirus (see Ty3/Gypsy family).
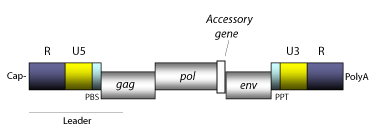
A canonical viral RNA genome consists of ~10Kb of RNA (+) with a 5'Cap and a 3'poly-A tail (Wang et al. 1975; Coffin and Billeter 1976).
In 5'-3' direction the viral RNA is structured as follows:
- A 5'direct repeat (R) of 18-250 nt
- A non-coding region of 75-250 nt (U5) that corresponds to the first portion of the retrotranscribed genome.
- A Primer Binding Site (PBS) of 18 nt, complementary to a specific zone of the 3' end of a tRNA normally provided by the host cell to start the retrotranscription.
- Open Reading Frames (ORFs) for gag, pol, and env genes and other accessory genes (in the case of complex retroviruses).
- A small region of ~10 A/G "Polypurine Tract" (PPT), responsible for starting the synthesis of the proviral (+) DNA strand.
- A non-coding zone of 200-1.200 nt (U3) containing the zone of promoters and constituting the 5' end of the proviral DNA.
- A 3'direct repeat (R) of 18-250 nt.
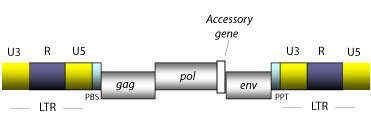
The retroviral RNA presents the same structural phenotype than cellular mRNAs. Its function, however, differs from that performed by cellular mRNAs in that following the infecting process, a retrovirus retrotranscribes itself in order to be inserted in the host cell as a proviral genome of DNA.
Prior to the integration, in the transcribing process the U3 zone is reordered upstream to the R and U5 zones generating a new LTR zone (U3/R/U5), which is subsequently duplicated (see LTRs formative process).
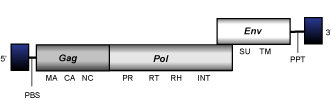
The gag gene codifies for a gag polyprotein containing the matrix (MA), capsid (CA) and the nucleocapsid (NC) domains which, in the maturation process, are spliced into independent peptides.
The pol gene codifies for a pol polyprotein containing the protease (PR), reverse transcriptase/ribonuclease H (RT/RNaseH), and integrase (INT) domains. However, PR may also be encoded by a gene alone, as a part of the gag polyprotein, or in frame with a dUTPase domain.
The env gene codifies for the envelope (env) glycoprotein, which in the maturation process is spliced into the outer surface (SU) membrane protein (the main antigen of the viral envelope), and the transmembrane (TM) protein.
Morphological types:
- Type A particles are unenveloped particles observed only intracellularly; they are most likely noninfectious products of the expression of endogenous retrovirus-like elements.
- Type B particles are enveloped extracellular particles characterized by presenting prominent envelope spikes and an acentric core.
- Type C particles are enveloped extracellular particles characterized by presenting barely visible spikes and a central core.
- Type D particles are enveloped extracellular particles, slightly larger than the other ones and characterized by presenting less prominent spikes.
Complexity:
Vertebrate retroviruses may be divided in simple and complex retroviruses. The main difference between them consists in that while simple retroviruses present the basal LTR-gag-pol-env-LTR genomic structure, complex retroviruses incorporate in their genomes some additional accessory genes usually needed to adjust diverse aspects of their replication and infectivity.
Strategy of transmission:
Based on the viral strategy of transmission, retroviruses that enter the germ lines and are vertically transmitted are referred to Endogenous Retroviruses (ERVs), to distinguish them from horizontally transmitted exogenous retroviruses (for a review in this topic see Gifford and Tristem 2003).
Evolutionary history and viral taxonomy:
Consistent with International Committee on the Taxonomy of Viruses (ICTV) (Fauquet et al. 2005), the inferred phylogeny of Retroviridae shows seven genera – Alpha-, Beta-, Gamma-, Delta-, Epsilon-, Spumaretrovirus and Lentivirus– that together with ERV-L elements we divide into three classes 1, 2 and 3 (according to Wilkinson et al. 1994;International Human Genome consortium 2001; 2002; Gifford and Tristem 2003; Gifford et al. 2005; Llorens et al. 2008). Class 1 comprises gamma- and epsilonretroviruses; class 2 includes lentiviruses, delta-, alpha- and betaretroviruses; and class 3 encompasses spumaretroviruses and ERV-L elements. Our database also includes an element called Snakehead retrovirus (SnRV) (Hart et al. 1996), which has unclear classification but that the common LTR retroelement phylogeny places within Class 1 (see, Llorens et al. 2009).
In differentiating the Retroviridae in the three classes 1, 2, and 3, comparative phylogenetic and network analyses reveal a network of relationships whereby class 1 can be related with Ty3/Gypsy lineages of plants and fungi -such as Tat and Athila elements and/or chromoviruses-, class 2 can be related to other Ty3/Gypsy lineages of insects, such as Micropia/Mdg3 clade; and class 3 with errantiviruses. In light of this polyphyletic scenario we proposed the three kings hypothesis Llorens et al. 2008, according to which the three Retroviridae classes can potentially be tracing three Ty3/Gypsy ancestors, emerged at different evolutionary times (for more details, see Llorens et al. 2009).
| Class | Genus | Example | Morphology |
|---|---|---|---|
| Class 1 | Gammaretrovirus | Murine Leukemia Virus | C-Type |
| Class 1 | Epsilonretrovirus | Walley Dermal Sarcoma Virus | C-Type |
| Class 2 | Alpharetroviridae | Avian Sarcoma and Leukosis Virus | C-Type |
| Class 2 | Betaretrovirus | Mouse Mammary Tumor Virus | B- and D-Type |
| Class 2 | Deltaretrovirus | Bovine Leukemia Virus | C-Type |
| Class 2 | Lentivirus | Human Immunodeficiency Virus | Cone-shaped core |
| Class 3 | Spumaretrovirus | Human Spumaretrovirus | C-Type |
| Class 3 | ERV-L | Murine Endogenous Retrovirus-Leucine | A-Type |
Class 1
This class is probably the most ancient branch of vertebrate retroviruses and comprises both gamma- and epsilonretroviruses.
Gammaretrovirus
The genus Gammaretrovirus collects both class 1 endogenous and exogenous C-type retroviruses to which phylogenetic analyses suggest RTVL-I-like viruses and epsilonretroviruses are related. Gammaretroviruses have a simple genome organization consisting in an internal region of 8-9 Kb in size, flanked by LTRs of 0.5 Kb and containing a PBS, ORFs for the gag, pol, and env genes typically observed in retroviruses, and a PPT adjacent to the 3'LTR.
Epsilonretrovirus
Epsilonretroviruses describe the C-type class 1 retroviruses of fishes. This genus is represented in this database by the Walleye Dermal Sarcoma Virus (WDSV) sequence (Martineau et al. 1991). Phylogenetic analyses suggest that epsilonretroviruses are closely related to gammaretroviruses. However, in difference to gammaretroviruses, WDSV is a complex retrovirus consisting in a full-length genome of 11 Kb in size, flanked by LTRs of 0,9 Kb in size and containing gag, pol, and env genes, a specific set of accessory genes, and also a PPT adjacent to the 3'LTR.
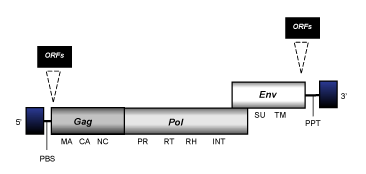
(figure not to scale)
Class 2
This class includes alpha-, beta- and deltaretroviruses and lentiviruses.
Alpharetrovirus
Alpharetroviruses are C-type class 2 retroviruses comprising both simple and complex retroviruses. Phylogenetic analyses suggest that alpharetroviruses are closely related to betaretroviruses. Alpharetroviruses present an internal region of approximately 6.8-9 Kb in size, flanked by LTRs of 0.3 Kb and characterized by the presence of a PBS, the usual gag, pol and env genes of retroviruses, a PPT adjacent to the 3'LTR, and in certain cases one or more accessory genes.
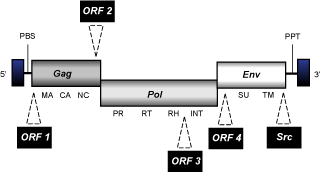
(figure not to scale)
Betaretrovirus
The genus Betaretrovirus comprises certain class 2 lineages of exogenous and endogenous B- and D-type retroviruses that normally present an internal region of approximately 7.5-9 Kb in size, flanked by LTRs of 0.3-1 Kb, and characterized by the presence of a PBS, a gag, dUTPase/pro, pol, and env plus OrfX genes organization, a PPT adjacent to the 3'LTR, and in certain cases one or more accessory genes.
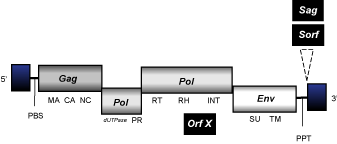
(figure not to scale)
Deltaretrovirus
Deltaretroviruses consist in a group of complex C-type class 2 retroviruses that constitute a well supported cluster along with Alpha- and betaretroviruses. Deltaretroviruses present an internal region of approximately 8-9 Kb in size, flanked by LTRs of 0.7-1 Kb and characterized by the presence of PBS, ORFs for the usual gag, pol, and env genes of retroviruses, a set of specific accessory genes, and also a PPT adjacent to the 3'LTR.
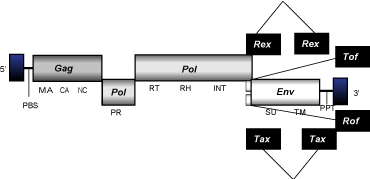
Lentivirus
Lentiviruses are complex class 2 retroviruses that present an internal region of approximately 8-9 Kb in size, flanked by LTRs of 0,2-1 Kb and characterized by the presence of a PBS, gag, pol, and env genes, and at least a PPT normally found adjacent to the 3'LTR. Lentiviruses also codify for a certain number of accessory genes common among all lentiviruses or, in certain cases, exclusively found depending of a clade, a genus, or a particular lentiviral species.
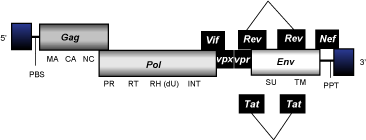
Class 3
Class 3 encompasses spumaretroviruses and ERV-L elements
Spumaretrovirus
Spumaretroviruses are complex C-type class 3 retroviruses whose internal region is approximately 13 Kb in size, flanked by LTRs of 1.7 Kb and characterized by the presence of a PBS, gag, pol and env ORFs, a specific set of accessory genes, and a PPT adjacent to the 3'LTR.
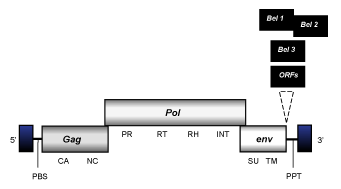
(figure not to scale)
ERV-L
ERV-L particles are A-type class 3 non-enveloped endogenous elements providing a putative evolutionary intermediate between classical intracellular retrotransposons and infectious retroviruses (Benit et al. 1999). ERV-L elements are represented in this database by the Murine Endogenous Retrovirus-Leucine (MuERV-L) sequence (Benit et al. 1997). The internal region of this element is 6 Kb in size, flanked by LTRs of 0.5 Kb, and characterized by the presence of a PBS, two ORFs for the gag, and pol genes typically observed in other LTR retrotransposons and retroviruses, as well as an additional dUTPase gene downstream to the integrase region, similar to that observed in betaretroviruses and non-primate lentiviruses, and a PPT normally adjacent to the 3'LTR (for more details see MuERV-L).